When imagining the future of technology, sometimes all we need to do is look out the window — or into a microscope.
Our researchers take inspiration from nature to redefine what a computer can be, from data storage using synthetic DNA, to sensors modeled on insects and leaves. We also advance technologies to help solve biology’s biggest mysteries, such as computational approaches for understanding the mechanisms of disease and brain-computer interfaces that can restore or augment physical function and mobility.
Research Groups & Labs
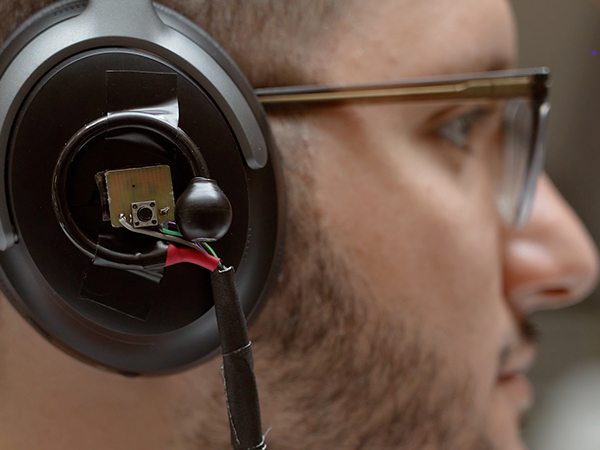

Mobile Intelligence Lab
The interdisciplinary Mobile Intelligence Lab builds intelligent systems and tools for tackling hard technical and societal problems, including battery-free computing, medical diagnostics, augmented human perception and more.
Systems Neuroscience & AI Lab (SNAIL)
SNAIL develops computational models and algorithms for understanding how single-trial neural population activity drives our abilities to generate movements, make decisions, and learn from experience.
Allen School Faculty
Assistant Professor
Professor
Assistant Professor
Associate Professor
Centers & Initiatives
The Institute for Medical Data Science (IMDS) is a joint effort among the Schools of Medicine and Public Health and the College of Engineering, including the Allen School to lead the development and implementation of cutting-edge AI and data science methods in medical data science. By harnessing the power of AI across diverse health determinants, IMDS aims to improve patient health, provider satisfaction, and healthcare operations, particularly in the Pacific Northwest region.
The Community-Engaged Computing Initiative (CECI) is a joint initiative to support community-centered scholarship and research within the broad computing and information field. Co-led by the Paul G. Allen School of Computer Science & Engineering, the Department of Human Centered Design & Engineering, and the Information School, the initiative was launched in 2025 through a gift from Google. CECI supports projects that bring UW faculty and graduate students together with community partners to bring sustainable, equitable, and inclusive technology into real-world contexts.
Highlights
Allen School News
American Academy of Arts & Sciences
GeekWire









